Input-output & lazy-loading#
Pynapple provides multiple ways to load data. The two main formats are NWB and NPZ. In addition, raw data can be loaded through the NEO library.
Each pynapple objects can be saved as a npz with a Pynapple-specific structure and loaded as a npz.
In addition, the Folder class helps you walk through a set of nested folders to load/save npz/nwb files.
NWB#
When loading a NWB file, pynapple will walk through it and test the compatibility of each data structure with a
pynapple objects. If the data structure is incompatible, pynapple will ignore it. The class that deals with reading
NWB file is nap.NWBFile. You can pass the path to a NWB file or directly an opened NWB file. Alternatively
you can use the function nap.load_file.
Note
Creating the NWB file is outside the scope of pynapple. The NWB files used here have already been created before. Multiple tools exists to create NWB file automatically. You can check neuroconv, NWBGuide or even NWBmatic.
data = nap.load_file(nwb_path)
print(data)
A2929-200711
┍━━━━━━━━━━━━━━━━━━━━━━━┯━━━━━━━━━━━━━┑
│ Keys │ Type │
┝━━━━━━━━━━━━━━━━━━━━━━━┿━━━━━━━━━━━━━┥
│ units │ TsGroup │
│ position_time_support │ IntervalSet │
│ epochs │ IntervalSet │
│ z │ Tsd │
│ y │ Tsd │
│ x │ Tsd │
│ rz │ Tsd │
│ ry │ Tsd │
│ rx │ Tsd │
┕━━━━━━━━━━━━━━━━━━━━━━━┷━━━━━━━━━━━━━┙
Pynapple will give you a table with all the entries of the NWB file that are compatible with a pynapple object.
When parsing the NWB file, nothing is loaded. The NWBFile class keeps track of the position of the data within the NWB file with a key. You can see it with the attributes key_to_id.
data.key_to_id
{'units': 'de078912-a093-4e6b-b23c-8647885a7188',
'position_time_support': '69ab7512-2d0a-45ba-af61-fae750c37f53',
'epochs': '2cca3914-ede4-41b8-b500-502465cbed95',
'z': 'e2142cf8-59d9-4637-b55d-796871c38e47',
'y': 'c5a8bba7-2699-4450-aae1-6f5a05309828',
'x': '2ba4ba5e-c6dc-4b9c-b69c-d0763240df46',
'rz': '682f8edf-91cb-4c15-bdf2-5cc5327803f0',
'ry': '57874425-e60e-4fe1-8bd3-7cb06d2adbfc',
'rx': '48597919-cce7-4724-9f72-db6e164c3c3f'}
Loading an entry will get pynapple to read the data.
z = data['z']
print(data['z'])
Time (s)
---------- ---------
670.6407 -0.195725
670.649 -0.19511
670.65735 -0.194674
670.66565 -0.194342
670.674 -0.194059
670.68235 -0.193886
670.69065 -0.193676
...
1199.94495 0.000398
1199.95325 -0.000552
1199.9616 -0.001479
1199.96995 -0.00237
1199.97825 -0.003156
1199.9866 -0.003821
1199.99495 -0.004435
dtype: float64, shape: (63527,)
Internally, the NWBClass has replaced the pointer to the data with the actual data.
While it looks like pynapple has loaded the data, in fact it did not. By default, calling the NWB object will return an HDF5 dataset.
print(type(z.values))
<class 'h5py._hl.dataset.Dataset'>
Notice that the time array is always loaded.
print(type(z.index.values))
<class 'numpy.ndarray'>
This is very useful in the case of large dataset that do not fit in memory. You can then get a chunk of the data that will actually be loaded.
z_chunk = z.get(670, 680) # getting 10s of data.
print(z_chunk)
Time (s)
---------- ---------
670.6407 -0.195725
670.649 -0.19511
670.65735 -0.194674
670.66565 -0.194342
670.674 -0.194059
670.68235 -0.193886
670.69065 -0.193676
...
679.9485 0.062836
679.95685 0.062831
679.96515 0.062789
679.9735 0.062756
679.98185 0.06277
679.99015 0.062819
679.9985 0.062878
dtype: float64, shape: (1124,)
Data are now loaded.
print(type(z_chunk.values))
<class 'numpy.ndarray'>
You can still apply any high level function of pynapple. For example here, we compute some tuning curves without preloading the dataset.
tc = nap.compute_tuning_curves(data['units'], data['y'], 10)
Warning
Carefulness should still apply when calling any pynapple function on a memory map. Pynapple does not implement any batching function internally. Calling a high level function of pynapple on a dataset that do not fit in memory will likely cause a memory error.
To change this behavior, you can pass lazy_loading=False when instantiating the NWBClass.
data = nap.NWBFile(nwb_path, lazy_loading=False)
z = data['z']
print(type(z.d))
<class 'numpy.ndarray'>
Raw data loading & NEO compatibility#
Raw data can be loaded with pynapple through the NEO library.
Internally, pynapple uses the NEO raw IO classes to read the data and convert them to one of the pynapple time series objects.
This is done lazily through the nap.EphysReader class.
See here the list of supported formats of python-neo.
The minimal example to load a dataset is to instantiate the EphysReader class with the path to the data file.
import pynapple as nap
data = nap.EphysReader("path_to_your_file")
Let us look at a couple of examples where we load LFP and spike data from different formats.
The EphysReader class will automatically detect the format of the data and load it accordingly.
To help the detection, you can also pass the format of the data with the argument format.
The format should be one of the supported formats of NEO. For example, if you have a binary file recorded with Neuroscope, you can pass format="NeuroscopeIO".
Neuroscope / Binary file#
If you have a session recorded as binary files with Neuroscope, you can load it with the EphysReader class or directly
with the function nap.load_binary_file.
📂 my_session
┣ 📄 my_session.dat
┗ 📄 my_session.xml
>>> import pynapple as nap
>>> data = nap.EphysReader("path/my_session", format="NeuroscopeIO")
my_session
┍━━━━━━━━━━━━━━━━━━━━━┯━━━━━━━━━━┑
│ Key │ Type │
┝━━━━━━━━━━━━━━━━━━━━━┿━━━━━━━━━━┥
│ my_session.dat │ TsdFrame │
┕━━━━━━━━━━━━━━━━━━━━━┷━━━━━━━━━━┙
>>> data["my_session.dat"]
Time (s) 0 1 2 3 4 ...
---------- --------- --------- --------- --------- --------- -----
0.0 0.274353 0.249023 0.3125 0.17395 0.240173 ...
5e-05 0.262146 0.205078 0.325928 0.163574 0.235901 ...
0.0001 0.263062 0.221863 0.325317 0.181885 0.266113 ...
... ...
1199.99585 -0.181885 -0.186157 -0.283508 -0.15686 -0.209961 ...
1199.9959 -0.098877 -0.209961 -0.291138 -0.213623 -0.209656 ...
1199.99595 -0.109863 -0.141296 -0.257874 -0.188293 -0.163879 ...
dtype: int16, shape: (23999920, 16)
A more direct way to load binary file is to use the function nap.load_binary_file which will return a TsdFrame object directly.
>>> import pynapple as nap
>>> data = nap.load_binary_file("path/my_session/my_session.dat",
channel=None # A list of channels to return. If None, returns all channels.
n_channels=16, # The number of channels in the binary file. Only used if channel is None.
sampling_rate=20000, # The sampling rate of the data.
precision='int16' # The precision of the data. Can be 'int16', 'float32' or 'float64'.
bytes_size=2 # The byte size of the data. Can be 2, 4 or 8.
)
>>> data
Time (s) 0 1 2 3 4 ...
---------- --------- --------- --------- --------- --------- -----
0.0 0.274353 0.249023 0.3125 0.17395 0.240173 ...
5e-05 0.262146 0.205078 0.325928 0.163574 0.235901 ...
0.0001 0.263062 0.221863 0.325317 0.181885 0.266113 ...
... ...
1199.99585 -0.181885 -0.186157 -0.283508 -0.15686 -0.209961 ...
1199.9959 -0.098877 -0.209961 -0.291138 -0.213623 -0.209656 ...
1199.99595 -0.109863 -0.141296 -0.257874 -0.188293 -0.163879 ...
dtype: float32, shape: (23999920, n_channels)
OpenEphys#
A typical OpenEphys dataset has the following structure:
📂 Record Node 109
┣ 📄 settings.xml
┗ 📂 experiment1
┗ 📂 recording1
┣ 📄 structure.oebin
┣ 📄 sync_messages.txt
┣ 📂 continuous
│ ┗ 📂 Neuropix-PXI-103.ProbeA
│ ┣ 📄 continuous.dat
│ ┣ 📄 sample_numbers.npy
│ ┗ 📄 timestamps.npy
┗ 📂 events
┣ 📂 MessageCenter
│ ┣ 📄 sample_numbers.npy
│ ┣ 📄 text.npy
│ ┗ 📄 timestamps.npy
┗ 📂 Neuropix-PXI-103.ProbeA
┗ 📂 TTL
┣ 📄 full_words.npy
┣ 📄 sample_numbers.npy
┣ 📄 states.npy
┗ 📄 timestamps.npy
This is a recording from a Neuropixel 2.0 probe, which does not have dedicated LFP channels.
Instead, we can visualize the raw data directly, using EphysReader as follows:
>>> import pynapple as nap
>>> data = nap.EphysReader("path/to/Record Node 109", format="OpenEphysBinaryIO")
Record Node 109
┍━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━┯━━━━━━━━━━┑
│ Key │ Type │
┝━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━┿━━━━━━━━━━┥
│ 0:Record Node 109#Neuropix-PXI-103.ProbeA │ TsdFrame │
│ 1:Record Node 109#Neuropix-PXI-103.ProbeASYNC │ TsdFrame │
│ 0:Messages │ Ts │
│ 1:Neuropixels PXI Sync │ Ts │
┕━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━┷━━━━━━━━━━┙
>>> data["0:Record Node 109#Neuropix-PXI-103.ProbeA"]
Time (s) 0 1 2 3 4 ...
------------ -------- ------- -------- -------- -------- -----
8.044366667 -138.06 -91.65 -115.245 -187.59 -130.455 ...
8.0444 -132.795 -82.485 -106.08 -185.445 -131.235 ...
8.044433333 -132.015 -90.87 -105.3 -187.59 -122.85 ...
... ...
68.606033333 33.54 56.55 50.31 -30.615 6.045 ...
68.606066667 35.88 56.55 45.045 -35.88 -1.56 ...
68.6061 32.76 61.815 51.87 -33.54 7.605 ...
dtype: int16, shape: (1816853, 384)
Plexon file#
If you have Plexon files, you can use EphysReader in the exact same way:
data = nap.EphysReader("File_plexon_3.plx", format="PlexonIO");
print(data)
File_plexon_3
┍━━━━━━━━━┯━━━━━━━━━━┑
│ Key │ Type │
┝━━━━━━━━━┿━━━━━━━━━━┥
│ 0:V │ TsdFrame │
│ TsGroup │ TsGroup │
┕━━━━━━━━━┷━━━━━━━━━━┙
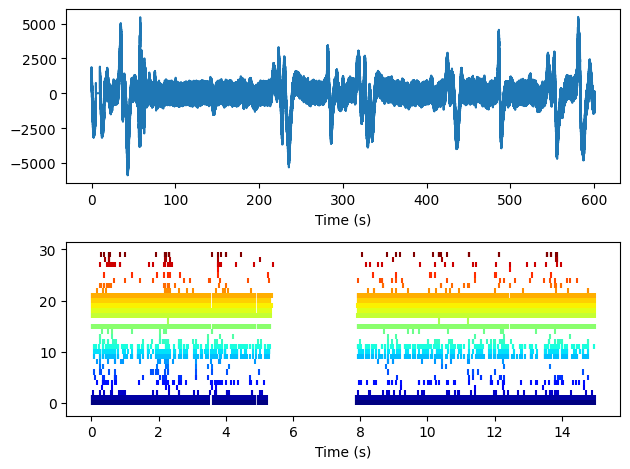
lfp = data['0:V']
spikes = data['TsGroup']
fig = plt.figure()
ax1 = fig.add_subplot(2, 1, 1)
ax2 = fig.add_subplot(2, 1, 2)
ax1.plot(lfp.t, lfp.d[:])
ax1.set_xlabel("Time (s)")
colors = plt.cm.jet(np.linspace(0, 1, len(spikes)))
ax2.eventplot([spikes[i].t for i in spikes.keys()], colors=colors)
ax2.set_xlabel("Time (s)")
plt.tight_layout()
plt.show()
Saving as NPZ#
Pynapple objects have save methods to save them as npz files.
tsd = nap.Tsd(t=np.arange(10), d=np.arange(10))
tsd.save("my_tsd.npz")
print(nap.load_file("my_tsd.npz"))
Time (s)
---------- --
0 0
1 1
2 2
3 3
4 4
5 5
6 6
7 7
8 8
9 9
dtype: int64, shape: (10,)
To load a NPZ to pynapple, it must contain particular set of keys.
print(np.load("my_tsd.npz"))
NpzFile 'my_tsd.npz' with keys: t, d, start, end, type
When the pynapple object have metadata, they are added to the NPZ file.
tsgroup = nap.TsGroup({
0:nap.Ts(t=[0,1,2]),
1:nap.Ts(t=[0,1,2])
}, metadata={"my_label":["a", "b"]})
tsgroup.save("group")
print(np.load("group.npz", allow_pickle=True))
NpzFile 'group.npz' with keys: type, _metadata, t, index, keys...
By default, they are added within the _metadata key:
print(dict(np.load("group.npz", allow_pickle=True))["_metadata"])
{'my_label': array(['a', 'b'], dtype='<U1')}
Memory map#
Numpy memory map#
Pynapple can work with numpy.memmap.
print(type(data))
<class 'numpy.memmap'>
Instantiating a pynapple TsdFrame will keep the data as a memory map.
eeg = nap.TsdFrame(t=timestep, d=data)
print(eeg)
Time (s) 0 1 2
---------- --------- --------- ----------
0 -0.418965 -1.87181 0.45505
1 -1.73578 0.272019 1.28516
2 1.03285 0.199289 -1.02125
3 0.347712 0.617409 -0.0501544
4 -0.739145 1.18075 -0.741286
5 -0.352571 -0.725824 0.589858
6 0.961491 -0.472428 0.427632
7 0.505873 0.600311 0.899022
8 -1.79447 1.00871 -1.48126
9 1.32282 -0.285034 1.02818
dtype: float32, shape: (10, 3)
We can check the type of eeg.values.
print(type(eeg.values))
<class 'numpy.memmap'>
Zarr#
It is possible to use Higher level library like zarr also not directly.
import zarr
zarr_array = zarr.zeros((10000, 5), chunks=(1000, 5), dtype='i4')
timestep = np.arange(zarr_array.shape[0])
tsdframe = nap.TsdFrame(t=timestep, d=zarr_array)
/home/runner/.local/lib/python3.12/site-packages/pynapple/core/utils.py:198: UserWarning: Converting 'd' to numpy.array. The provided array was of type 'Array'.
warnings.warn(
As the warning suggest, zarr_array is converted to numpy array.
print(type(tsdframe.d))
<class 'numpy.ndarray'>
To maintain a zarr array, you can change the argument load_array to False.
tsdframe = nap.TsdFrame(t=timestep, d=zarr_array, load_array=False)
print(type(tsdframe.d))
<class 'zarr.core.Array'>
Within pynapple, numpy memory map are recognized as numpy array while zarr array are not.
print(type(data), "Is np.ndarray? ", isinstance(data, np.ndarray))
print(type(zarr_array), "Is np.ndarray? ", isinstance(zarr_array, np.ndarray))
<class 'numpy.memmap'> Is np.ndarray? True
<class 'zarr.core.Array'> Is np.ndarray? False
Similar to numpy memory map, you can use pynapple functions directly.
ep = nap.IntervalSet(0, 10)
tsdframe.restrict(ep)
Time (s) 0 1 2 3 4
---------- --- --- --- --- ---
0 0 0 0 0 0
1 0 0 0 0 0
2 0 0 0 0 0
3 0 0 0 0 0
4 0 0 0 0 0
5 0 0 0 0 0
6 0 0 0 0 0
7 0 0 0 0 0
8 0 0 0 0 0
9 0 0 0 0 0
10 0 0 0 0 0
dtype: int32, shape: (11, 5)
group = nap.TsGroup({0:nap.Ts(t=[10, 20, 30])})
sta = nap.compute_event_triggered_average(tsdframe, group, 1, (-2, 3))
print(type(tsdframe.values))
print("\n")
print(sta)
<class 'zarr.core.Array'>
Time (s)
---------- -----------------
-2 [[0. ... 0.] ...]
-1 [[0. ... 0.] ...]
0 [[0. ... 0.] ...]
1 [[0. ... 0.] ...]
2 [[0. ... 0.] ...]
3 [[0. ... 0.] ...]
dtype: float64, shape: (6, 1, 5)
