Correlograms & ISI#
Let’s generate some data. Here we have two neurons recorded together. We can group them in a TsGroup.
ts1 = nap.Ts(t=np.sort(np.random.uniform(0, 1000, 2000)), time_units="s")
ts2 = nap.Ts(t=np.sort(np.random.uniform(0, 1000, 1000)), time_units="s")
epoch = nap.IntervalSet(start=0, end=1000, time_units="s")
ts_group = nap.TsGroup({0: ts1, 1: ts2}, time_support=epoch)
print(ts_group)
Index rate
------- ------
0 2
1 1
Autocorrelograms#
We can compute their autocorrelograms meaning the number of spikes of a neuron observed in a time windows centered around its own spikes.
For this we can use the function compute_autocorrelogram.
We need to specifiy the binsize and windowsize to bin the spike train.
autocorrs = nap.compute_autocorrelogram(
group=ts_group, binsize=100, windowsize=1000, time_units="ms", ep=epoch # ms
)
print(autocorrs)
0 1
-0.9 0.9650 0.95
-0.8 1.0775 0.98
-0.7 0.9000 0.94
-0.6 1.0375 0.99
-0.5 0.9675 0.98
-0.4 1.0375 0.95
-0.3 0.9050 0.98
-0.2 1.0175 0.98
-0.1 1.0500 0.99
0.0 0.0000 0.00
0.1 1.0500 0.99
0.2 1.0175 0.98
0.3 0.9050 0.98
0.4 1.0375 0.95
0.5 0.9675 0.98
0.6 1.0375 0.99
0.7 0.9000 0.94
0.8 1.0775 0.98
0.9 0.9650 0.95
The variable autocorrs is a pandas DataFrame with the center of the bins
for the index and each column is an autocorrelogram of one unit in the TsGroup.
Cross-correlograms#
Cross-correlograms are computed between pairs of neurons.
crosscorrs = nap.compute_crosscorrelogram(
group=ts_group, binsize=100, windowsize=1000, time_units="ms" # ms
)
print(crosscorrs)
0
1
-0.9 1.025
-0.8 1.090
-0.7 1.030
-0.6 0.895
-0.5 1.030
-0.4 1.055
-0.3 1.000
-0.2 1.035
-0.1 1.075
0.0 0.985
0.1 0.950
0.2 1.090
0.3 1.080
0.4 0.975
0.5 0.930
0.6 1.070
0.7 0.955
0.8 0.995
0.9 1.080
Column name (0, 1) is read as cross-correlogram of neuron 0 and 1 with neuron 0 being the reference time.
Event-correlograms#
Event-correlograms count the number of event in the TsGroup based on an event timestamps object.
eventcorrs = nap.compute_eventcorrelogram(
group=ts_group, event = nap.Ts(t=[0, 10, 20]), binsize=0.1, windowsize=1
)
print(eventcorrs)
0 1
-0.9 1.801802 0.000000
-0.8 0.000000 0.000000
-0.7 5.405405 3.174603
-0.6 0.000000 0.000000
-0.5 3.603604 0.000000
-0.4 0.000000 0.000000
-0.3 1.801802 0.000000
-0.2 1.801802 3.174603
-0.1 0.000000 0.000000
0.0 0.000000 0.000000
0.1 1.801802 0.000000
0.2 0.000000 3.174603
0.3 0.000000 0.000000
0.4 1.801802 0.000000
0.5 1.801802 0.000000
0.6 0.000000 0.000000
0.7 0.000000 0.000000
0.8 0.000000 0.000000
0.9 0.000000 0.000000
Interspike interval (ISI) distribution#
The interspike interval distribution shows how the time differences between subsequent spikes (events) are distributed.
The input can be any object with timestamps. Passing epochs restricts the computation to the given epochs.
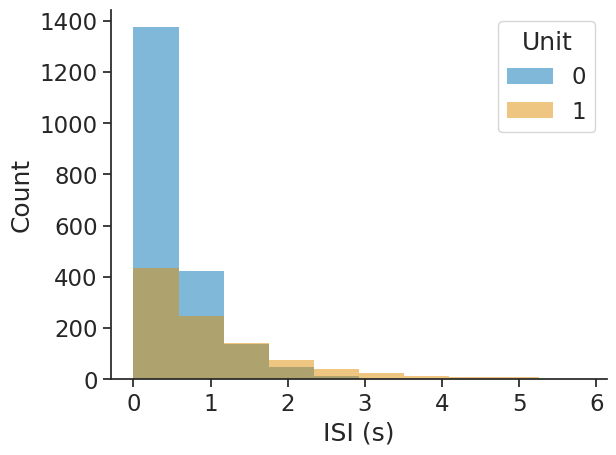
The output will be a dataframe with the bin centres as index and containing the corresponding ISI counts per unit.
isi_distribution = nap.compute_isi_distribution(
data=ts_group, bins=10, epochs=epoch
)
print(isi_distribution)
0 1
0.292762 1375 436
0.877336 424 248
1.461909 136 143
2.046483 49 76
2.631056 11 41
3.215630 2 25
3.800204 0 14
4.384777 0 8
4.969351 2 7
5.553924 0 1
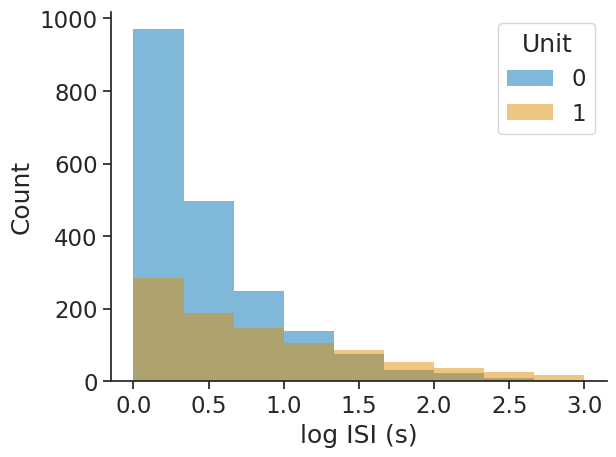
The bins argument allows for choosing either the number of bins as an integer or the bin edges as an array directly:
isi_distribution = nap.compute_isi_distribution(
data=ts_group, bins=np.linspace(0, 3, 10), epochs=epoch
)
print(isi_distribution)
0 1
0.166667 970 285
0.500000 498 189
0.833333 248 147
1.166667 139 105
1.500000 74 86
1.833333 31 54
2.166667 24 37
2.500000 8 25
2.833333 4 18
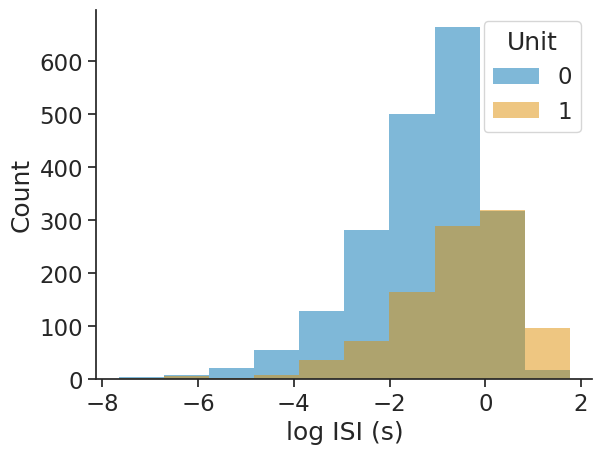
The log_scale argument allows for applying the log-transform to the ISIs:
isi_distribution = nap.compute_isi_distribution(
data=ts_group, bins=10, log_scale=True, epochs=epoch
)
print(isi_distribution)
0 1
-7.180847 5 2
-6.239096 8 6
-5.297344 21 3
-4.355592 55 8
-3.413840 129 36
-2.472089 281 73
-1.530337 500 165
-0.588585 664 289
0.353166 318 320
1.294918 18 97